Materials
Photo of Alex Jones in a University of Georgia (UGA) lab in 2015. While a doctoral student at UGA, Jones studied antibacterial bioplastics.
Materials
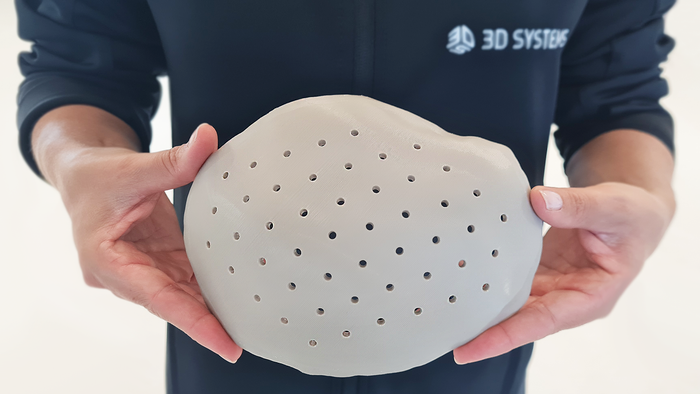
These Antibacterial Bioplastics Will Make You Say, ‘What the Cluck?’These Antibacterial Bioplastics Will Make You Say, ‘What the Cluck?’
Trivia Tuesday: Why did researchers take an interest in chicken eggs?
Sign up for the QMED & MD+DI Daily newsletter.













.png?width=300&auto=webp&quality=80&disable=upscale)

















.png?width=300&auto=webp&quality=80&disable=upscale)
