Materials
3D-printed PEEK cranial implant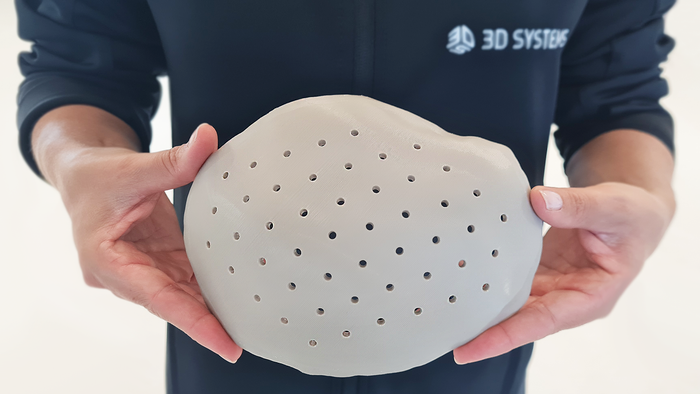
Implants
3D-printed PEEK-based Cranial Implants Cleared by FDA3D-printed PEEK-based Cranial Implants Cleared by FDA
The VSP PEEK Cranial Implant system includes a complete FDA-cleared workflow comprising segmentation and 3D-modeling software, a printer developed by 3D Systems, and PEEK resin from Evonik.
Sign up for the QMED & MD+DI Daily newsletter.









.png?width=300&auto=webp&quality=80&disable=upscale)